Bionanophotonic Systems laboratory
Prof. Hatice ALTUG
At the BIOnanophotonic Systems Laboratory, we develop ultra‑sensitive spectroscopy and sensing platforms for real‑time, label‑free, high‑throughput detection of trace biomolecules. Our work uses nanoplasmonics, metamaterials, and micro‑/nanofluidics to enable precise analyte trapping and manipulation. We also introduce new fabrication methods for low‑cost, large‑area, high‑throughput device production. In addition to biochemical sensing and spectroscopy, we investigate nanophotonic approaches for on‑chip optical communications.
Antanasijevic Lab – Virology and Structural Immunology
Prof. Aleksandar ANTANASIJEVIC
Our lab uses state-of-the-art electron microscopy to study antibody-antigen interactions in many different contexts, with the ultimate goal to define the molecular rules that govern them.
Laboratory of Protein and Cell Engineering
Prof. Patrick BARTH
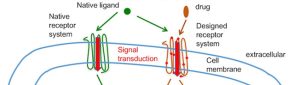
Our lab develops computational approaches to predict and design protein structures, dynamics and functions. We apply these tools to design therapeutic proteins and engineer cells with novel functions.
G-LAB UPCOURTINE
Prof. Grégoire COURTINE
We leverage the most advanced single-cell and spatial methodologies to chart the molecular landscape of spinal cord injury, stroke and neurodegenerative diseases. With these technologies, our goal is to establish molecularly informed strategies to achieve genetic re-engineering of damaged neural tissues.
Laboratory for Biomolecular Modeling
Prof. Matteo DAL PERARO
Integrative structural biology, molecular design and nanopore sensing.
Lipid Cell Biology Laboratory
Prof. Giovanni D’ANGELO
The functional properties of cell membranes strongly depend on their lipid composition and different membranes both within the cell and among different cells have different lipid composition. At the Lipid Cell Biology Lab, we aim at understanding the meaning of compositional variability in cell membranes by studying the mechanisms by which the lipid composition is determined.
Laboratory of Systems Biology and Genetics
Prof. Bart DEPLANCKE
The Laboratory of Systems Biology and Genetics (LSBG) studies genome organization, regulation, and variation through three pillars: “Adipo” explores mesenchymal stromal cell function in adipose biology, “Geno” examines how regulatory variation shapes diversity, and “Techo” develops advanced microfluidic, sequencing, and computational tools to drive discoveries in both areas.
Laboratory for Bio- and Nano- Instrumentation
Prof. Georg FANTNER
We develop new measurement tools and instruments to characterize and manipulate biological samples (molecules, cells, tissue) at the nanoscale. Our focus is to enable new biological research through advancing scanning probe techniques such as atomic force microscopy, scanning ion conductance microscopy, single molecule measurements and 3D nanoscale tomography.
Laboratory of Biophysical Chemistry of Macromolecules
Prof. Beat FIERZ
Our lab develops advanced single-molecule fluorescence and FRET microscopy methods to uncover the molecular mechanisms governing chromatin function, transcription factor dynamics and epigenetic regulation. We also pioneer chemical biology approaches to decode post-translational modifications in both chromatin and microtubules, revealing fundamental regulatory principles in genome organization and cellular architecture.

Gallini lab – Empowerment of healthy cells to prevent skin cancer
Prof. Sara GALLINI
Our lab studies how healthy skin stem cells control and suppress pre-cancerous mutant cells. We use in vivo imaging and molecular approaches to uncover the cellular and signaling mechanisms that preserve tissue integrity and prevent tumor initiation.
Gönczy Lab – Cell and Developmental Biology
Prof. Pierre GÖNCZY
We are interested in understanding fundamental cellular processes, including in the context of development. Our main focus is on the mechanisms governing centriole assembly, as well as asymmetric cell division.
Laboratory of Life Sciences Electronics
Prof. Carlotta GUIDUCCI
At the forefront of this innovation, our research focuses on miniaturizing analytical and processing workflows using cutting-edge microfluidics and microsystem technologies. Working at the intersection of fundamental research and applied bioanalytics, we are committed to bridging the gap between laboratory innovation and clinical application—developing tools that are not only technically advanced but also accessible, scalable, and impactful for real-world healthcare.
Laboratory of Computational Systems Biotechnology
Prof. Vassily HATZIMANIKATIS
At the Laboratory of Computational Systems Biotechnology (LCSB), we work at the interface of synthetic and systems biology to identify the design principles of biological processes for medical and biotechnological applications.
Our research areas of interest include: Cellular Networks, Kinetic Modelling, Novel Biotransformations.
Laboratory of Synthetic and Applied Microbiology
Prof. Markus JESCHEK
The LSAM develops synthetic microbes that sustainably produce biotechnological products ranging from bulk and speciality chemicals to proteins or entire cells. We combine ultrahigh-throughput experimental technology with cutting-edge machine-learning techniques to equip these microbes with new-to-nature functions and enable their design “à la carte”.
Mechanics of Soft and Biological Matter Laboratory
Prof. Sangwoo KIM
In the Mechanics of Soft and Biological Matter Laboratory (MESOBIO), we endeavor to gain a fundamental understanding of biological and living systems, as well as soft and active materials. By developing theoretical and computation frameworks based on physics and mechanics principles, we aim to understand how cellular and subcellular properties give rise to emergent architecture, dynamics, and mechanical states at the tissue level.

Kim Lab of Molecular and Synthetic Neuroengineering
Prof. Yoon KIM
We develop innovative bioengineering tools to study the brain while uncovering the molecular mechanisms that govern brain disease, signal transmission, and plasticity. By integrating protein engineering, structural biology, synthetic biology, and neurobiology, we bridge fundamental neuroscience with translational and therapeutic applications.
Laboratory of Brain Development and Biological Data Science
Prof. Gioele LA MANNO
We are a computational and developmental biology lab with a focus on understanding the complexities and dynamics of brain development. We combine single-cell, spatial omics technologies, bilogical data science and machine learning to discover how neural stem cells transform into “the misterious butterflies of the soul”.
Laboratory of Experimental Biophysics
Prof. Suliana MANLEY
We develop smart and super-resolution fluorescence microscopy methods. Our goals are to enable gentler live cell imaging, while adapting the measurement to the sample dynamics, and to correlate structure, dynamics, and function. We use these methods to study organelle dynamics, focusing on the mitochondrial life cycle.
Laboratory of Biomedical Microfluidics
Prof. Christoph MERTEN
The Laboratory for Biomedical Microfluidics (LBMM) develops new technologies for antibody discovery, immune repertoire analysis and personalized cancer therapy. Making use of assay miniaturization, a big focus is on processing limited patient material and performing single-cell analysis.
Laboratory of Computational and Systems Biology
Prof. Felix NAEF
Prof. Felix Naef leads the Computational Systems Biology Lab at EPFL. His team develops quantitative models and computational approaches to understand the dynamics of gene regulation and cellular rhythms. By integrating genomics, mathematical modeling, and biophysics, they investigate how biological clocks, circadian rhythms, and regulatory networks govern cellular behavior and maintain physiological balance.
Segmentation Timing and Dynamics Laboratory
Prof. Andy OATES
We study how a population of synchronized genetic oscillators – the segmentation clock – acts to segment the growing body axis of the vertebrate embryo. We use a combination of genetics, biochemistry, experimental embryology, fluorescence microscopy, image processing and computational modeling to understand the structure and dynamics of the zebrafish segmentation clock from molecular to tissue scales.
Microbial Mechanics lab
Prof. Alex PERSAT
The Persat lab uses an interdisciplinary bioengineering approach to investigate bacterial infections and the rise of antibiotic resistance. We combine tissue-engineered organoids with omics and imaging to decode the contributions of mechanics during bacterial infections and to discover novel therapeutic strategies to combat resistant pathogens.
Laboratory of the Physics of Biological Systems
Prof. Sahand RAHI
The Rahi lab works at the intersection of biophysics with systems and synthetic biology. We are interested in understanding computation and dynamics in genetic and cellular networks, and we develop new directed evolution approaches coupled to generative AI.
Living Patterns Laboratory
Prof. Guillermina RAMIREZ-SAN-JUAN
We are interested in understanding how biological patterning and function emerge from microscopic molecular interactions. Our primary focus is on the problems of flow generation by arrays of active filaments (cilia) and extreme cellular mechanics.
Molecular Spatial Omics Laboratory
Prof. Florian SCHUEDER
Our lab investigates how the spatial arrangement of biomolecules, from the molecular scale up to the mesoscale, drives biological function.By inventing and applying cutting-edge imaging hardware, DNA nanotechnology, and computational tools, we aim to reveal the principles that link molecular architecture to cellular and tissue-level behavior, enabling new insights across microbiology, systems biology, and beyond.
Laboratory of Biomaterials for Immunoengineering
Prof. Li TANG
Tang Laboratory is developing novel strategies to engineer the multi-dimensional immunity-disease interactions from various aspects, an emerging field called ‘immunoengineering’, in order to create safe and effective therapies against cancer and infectious diseases. Specifically, we leverage the power of metabolic and cellular bioengineering, synthetic chemistry and material engineering, and mechanical engineering to achieve controllable modulation of immune responses against diseases.
NeuroNA Chair in Epigenomics of Neurodevelopmental disorders – EpiGN
Prof. Fides ZENK
We engineer brain organoid systems to model early human development and uncover the molecular logic of brain formation. By developing and applying cutting-edge single-cell genomics technologies, we map how gene regulation and chromatin dynamics guide cell fate decisions.