Laboratory of Computational Science and Modeling COSMO
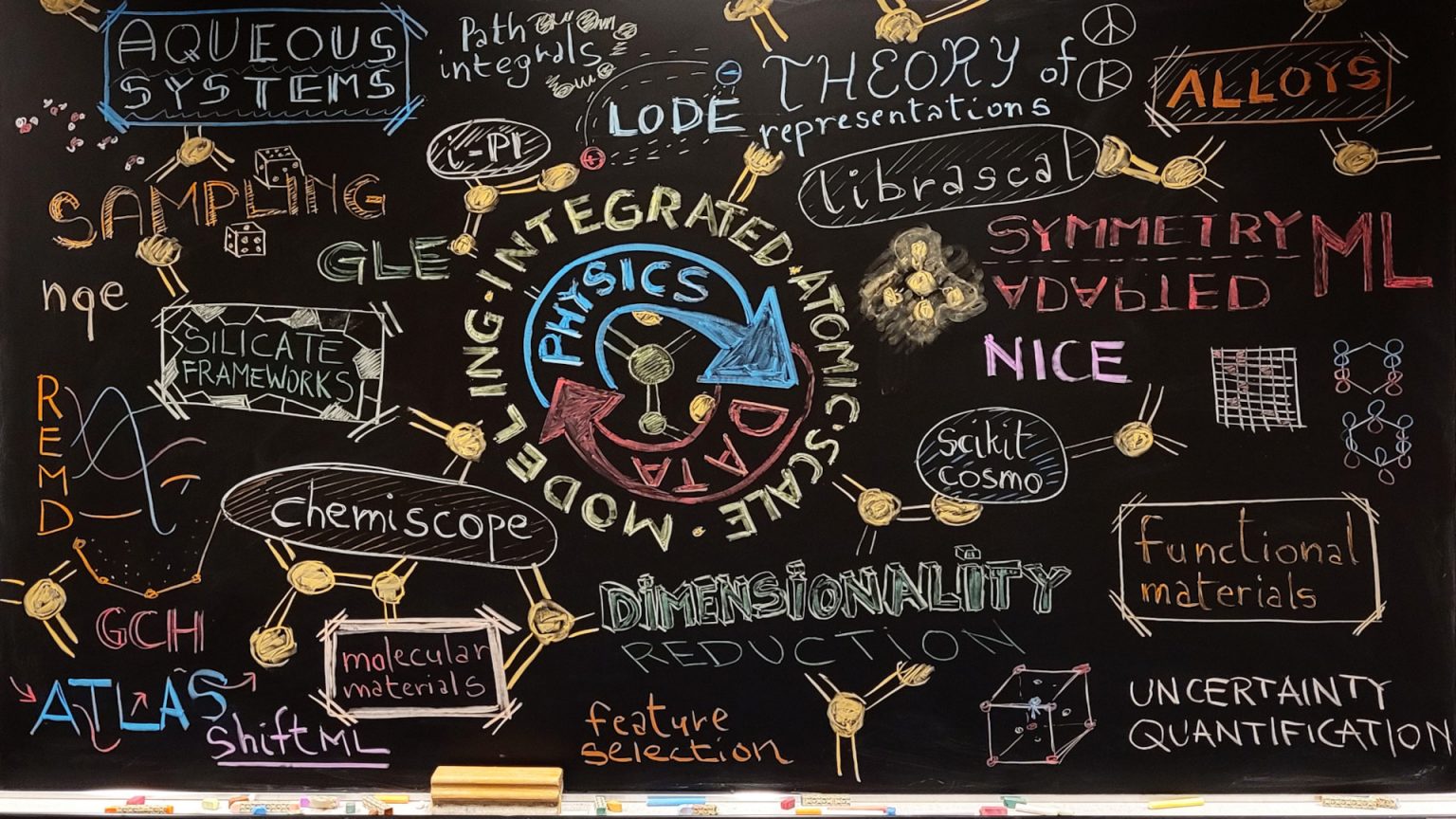
An overview of the activities of the lab, using physics and data-driven models to achieve predictive modeling of materials and molecules.
.
The macroscopic behaviour of materials is determined by the chemical and physical interactions between atoms, averaged over thermal and quantum mechanical fluctuations. At the laboratory of Computational Science and Modelling we simulate matter at the atomic scale, to be able to predict and optimise the properties of materials and molecules starting from fundamental physical principles.
To do so, we need to develop computer simulation methods that are both efficient and accurate, relying on an interdisciplinary expertise that combines understanding of statistical physics, quantum chemistry and materials engineering, with the mathematical and computational tools of data science and machine learning. We then apply these techniques to address materials science problems that are relevant for pressing societal issues – from the metallurgy of lightweight alloys for sustainable transportation, to the determination of the structure and behaviour of hydrogen-bonded materials such as pharmaceutical compounds and bio-inspired polymers.
You can learn more on what we do, and what methods we use, on the Research page, or on the News Archive. If you are interested in joining us, have a look at the Jobs page for open positions and application instructions.
Contacts
| Head of the laboratory: Prof. Michele Ceriotti tel: +41 (0)21 69 32939 | Administration: Ms. Isabel Nzazi tel: +41 (0)21 69 32925 | EPFL STI IMX COSMO MXG 338 (Bâtiment MXG) Station 12 CH-1015 Lausanne |
Links
Follow @labCOSMO Follow @ Mastodon View our code on github


