Although composed of discrete fat depots, adipose tissue forms a remarkably organized endocrine organ playing the major role in nutrient homeostasis, energy balance and immunity. Functionally two distinct types of fat are distinguished namely white and brown. White fat (WAT) is the primary site for energy storage and has been widely implicated in obesity and its associated pathologies such as insulin resistance, type 2 diabetes or more broadly, the metabolic syndrome. In contrast, the less abundant and cold-inducible brown fat (BAT) releases energy in form of heat through the highly specific expression of the uncoupling protein 1 (UCP1) (Ricquier 2011). Additionally, cells with brown-like characteristics, termed as beige/brite cells, arise within classically white adipose depots upon cold exposure (and other stimuli) in both rodents and humans. Typically, brown fat has been associated with a wide range of metabolically favourable indicators, leading to the broadly accepted view that activation of brown and beige fat tissues represents a promising strategy for solving global health challenges such as obesity and the metabolic syndrome.
Our group is unraveling the transcriptional mechanisms underlying the molecular basis of adipogenesis and cell type specific differentiation by applying integrative genomic approaches and single-cell transcriptomics as well as in vivo assays. Specifically, we:
-
-
Examine the origin, diversity and function of white, brite and brown fat cells using single cell transcriptomics
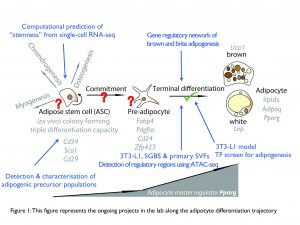
Using single-cell RNA sequencing (scRNA-seq), we aim to uncover the molecular identity of white, beige and brown fat cell precursors, map their location on adipocyte lineage trees, and determine cellular composition differences between fat depots both unperturbed and after intervention (i.e. exposure to cold to induce browning). Further, we will investigate the contribution of the different adipose cell precursors/populations to development of metabolism associated disorders such as obesity and metabolic syndrome.
-
Study gene regulatory networks mediating determination and terminal differentiation of white, brite and brown fat cells
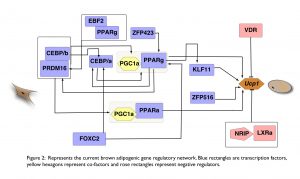
Although many of the key transcription factors controlling white fat cell differentiation have been characterized (Tang and Lane 2012), a cohesive model of the underlying molecular events has yet to be revealed. Further, the transcriptional regulation of brite and brown fat cell differentiation is substantially less well understood. Combining differential expression analyses (RNAseq/scRNA-seq), chromatin landscape data (histone modification ChIP-seq), as well as large-scale transcription factor (TF) perturbation screens, we aim to unravel regulatory networks that impact adipogenesis. Primarily, our ambitions in this field are focused on TFs (Gubelmann, Raghav and Schwalie et al., 2014) and their co-regulators (Raghav and Waszak et al., 2012). Ultimately, we pursue in-vivo validation of the identified factors in mouse models, in collaboration with Christian Wolfrum’s lab at ETH Zurich and Martin Klingenspor’s lab at TU Munich.
-
Probe the impact of genetic variation on adipogenic differentiation
In parallel to extensive analyses of murine gene regulatory networks underlying adipose cell differentiation and function we are focused on the molecular characterization of human adipogenic precursor populations. As an emerging and very promising source of autologous multipotent adult stem cells, adipose derived human mesenchymal stem cells (ASCs) are increasingly used in diverse biomedical applications. However, the lack of comprehensive molecular characterisation impedes its growing clinical implementation. Using the single cell transcription profiling, we are assessing the cellular heterogeneity of ASCs populations obtained from human lipoaspirate samples to reveal the traits associated with the highest multipotency. Another goal is to address the impact of genetic variation on differentiation capacities of ASCs (see the “Geno” Website Section). These projects are being done in collaboration with the Swiss Stem Cell Foundation (SSCF) in Lugano.
-
References
-
Ricquier, D. Uncoupling protein 1 of brown adipocytes, the only uncoupler: a historical perspective. Front. Endocrinol. (Lausanne) 2, 85 (2012).
-
S.K. Raghav*, S.M. Waszak*, I. Krier, A. Isakova, C. Gubelmann, T.S. Mikkelsen, and B. Deplancke. Integrative genomics identifies SMRT as a gatekeeper of adipogenesis through the transcription factors C/EBPb and KAISO, Molecular Cell, 46:335-350, 2012.
-
C. Gubelmann*, P.C. Schwalie*, S.K. Raghav*, E. Röder, T. Delessa, E. Kielmann, S.M. Waszak, A. Corsinotti, G. Udin, W. Holcombe, G. Rudofsky, D. Trono, C. Wolfrum, B. Deplancke. Identification of ZEB1 as a central component of the adipogenic gene regulatory network. eLife, 3:e03346, 2014.
-
Shapiro, E., T. Biezuner and S. Linnarsson (2013). “Single-cell sequencing-based technologies will revolutionize whole-organism science.” Nat Rev Genet 14(9): 618-630.
-
Tang, Q. and D. M. Lane (2012). “Adipogenesis: From Stem Cell to Adipocyte.” Annual review of biochemistry 81(1): 715-736.