Category: Highlights
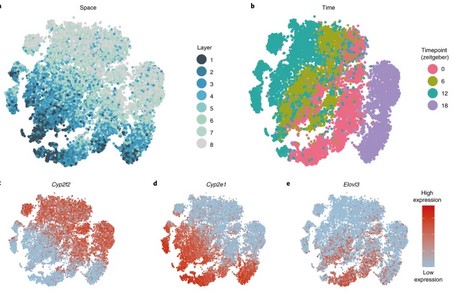
Space-time logic of liver gene expression at sub-lobular scale
The mammalian liver is a central hub for systemic metabolic homeostasis. Liver tissue is spatially structured, with hepatocytes operating in repeating lobules, and sub-lobule zones performing distinct functions. The liver is also subject to extensive temporal regulation, orchestrated by the interplay of the circadian clock, systemic signals and feeding rhythms. However, liver zonation has previously (…)
The circadian oscillator analysed at the single-transcript level
The circadian clock is an endogenous and self-sustained oscillator that anticipates daily environmental cycles. While rhythmic gene expression of circadian genes is well-described in populations of cells, the single-cell mRNA dynamics of multiple core clock genes remain largely unknown. Here we use single-molecule fluorescence in situ hybridisation (smFISH) at multiple time points to measure pairs (…)
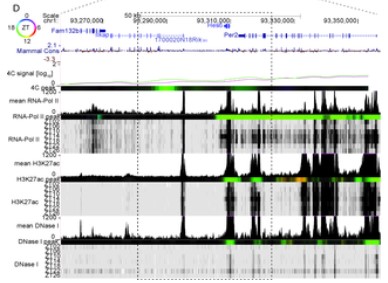
Oscillating and stable genome topologies underlie hepatic physiological rhythms during the circadian cycle
The circadian clock drives extensive temporal gene expression programs controlling daily changes in behavior and physiology. In mouse liver, transcription factors dynamics, chromatin modifications, and RNA Polymerase II (PolII) activity oscillate throughout the 24-hour (24h) day, regulating the rhythmic synthesis of thousands of transcripts. Also, 24h rhythms in gene promoter-enhancer chromatin looping accompany rhythmic mRNA (…)
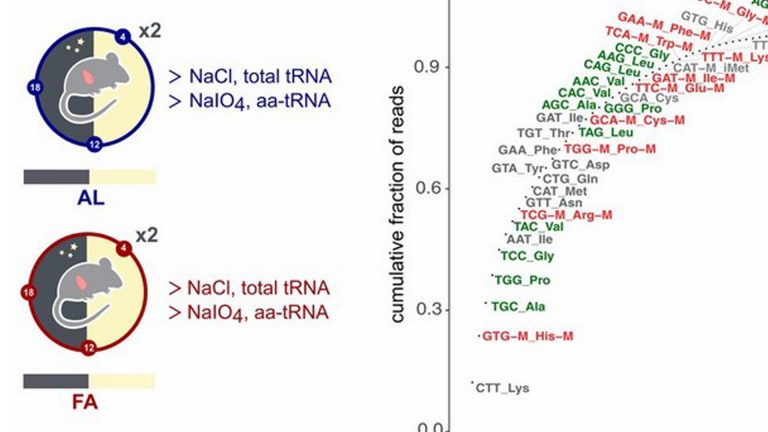
Robust landscapes of ribosome dwell times and aminoacyl-tRNAs in response to nutrient stress in liver
Translation depends on messenger RNA (mRNA)-specific initiation, elongation, and termination rates. While translation elongation is well studied in bacteria and yeast, less is known in higher eukaryotes. Here we combined ribosome and transfer RNA (tRNA) profiling to investigate the relations between translation elongation rates, (aminoacyl-) tRNA levels, and codon usage in mammals. We modeled codon-specific (…)

Modulation of transcriptional burst frequency by histone acetylation
Many mammalian genes are transcribed during short bursts of variable frequencies and sizes that substantially contribute to cell-to-cell variability. However, which molecular mechanisms determine bursting properties remains unclear. To probe putative mechanisms, we combined temporal analysis of transcription along the circadian cycle with multiple genomic reporter integrations, using both short-lived luciferase live microscopy and single-molecule (…)
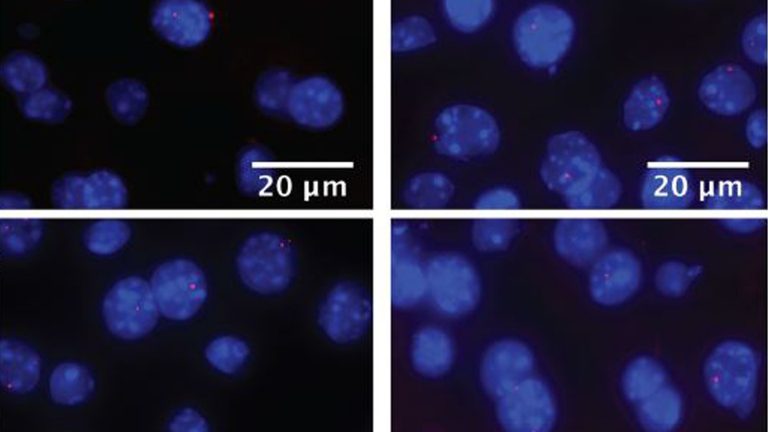
Clock-dependent chromatin topology modulates circadian transcription and behavior
The circadian clock in animals orchestrates widespread oscillatory gene expression programs, which underlie 24-h rhythms in behavior and physiology. Several studies have shown the possible roles of transcription factors and chromatin marks in controlling cyclic gene expression. However, how daily active enhancers modulate rhythmic gene transcription in mammalian tissues is not known. Using circular chromosome (…)
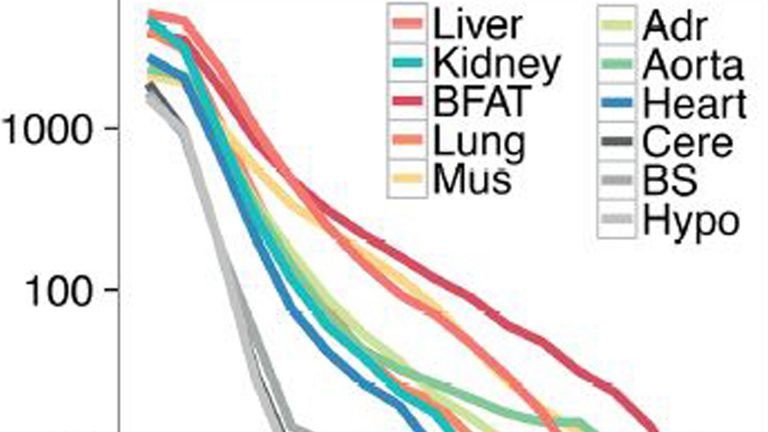
Transcription factor activity rhythms and tissue-specific chromatin interactions explain circadian gene expression across organs
Temporal control of physiology requires the interplay between gene networks involved in daily timekeeping and tissue function across different organs. How the circadian clock interweaves with tissue-specific transcriptional programs is poorly understood. Here, we dissected temporal and tissue-specific regulation at multiple gene regulatory layers by examining mouse tissues with an intact or disrupted clock over (…)