Despite extensive research and modelling, there is still a gap in our understanding of neurons — in particular in how they form networks. Using a bottom-up approach studying small-scale in vitro cultures, such as enforcing connections by physical constraints or quantifying self-assembled connections under electrical stimulation, we work towards building a solid understanding of universal neuronal behaviour. By creating setups that force specific, poorly understood situations to occur — such as axo-axonic connections — as well as reliably obtaining statistical trends of networks of limited size, we aim to build a stronger rule-set on which emergence occurs.
Spike Softening: a computationally efficient method for extracting functional connectivity maps from neuronal networks
The development of High-Density MicroElectrode Arrays (HD-MEAs) enables unprecedented simultaneous recording of thousands of individual neurons, raising strong interest in optimised analysis techniques — particularly for real-time applications such as closed-loop stimulation. A common characteristic derived from network recordings is the functional connectivity map, typically obtained via cross-correlation on each neuron pair. However, cross-correlation scales poorly with the number of samples, making it too slow for real-time HD-MEA deployment.
We propose an alternative algorithm called Spike Softening, inspired by Spike-Timing-Dependent Plasticity (STDP). The method successively approximates each neuron pair’s connection strength, ultimately converging to a functional connectivity map comparable to cross-correlation — but at significantly lower algorithmic cost. For each incoming Boolean spike array, an exponentially decaying tail is appended after each detected spike. At each neuron spike event, the tail value of all other neurons is accumulated into their pairwise weights, effectively identifying nodes that are time-correlated. The exponential base is a tunable parameter analogous to bin-width in cross-correlation, without its computational overhead.
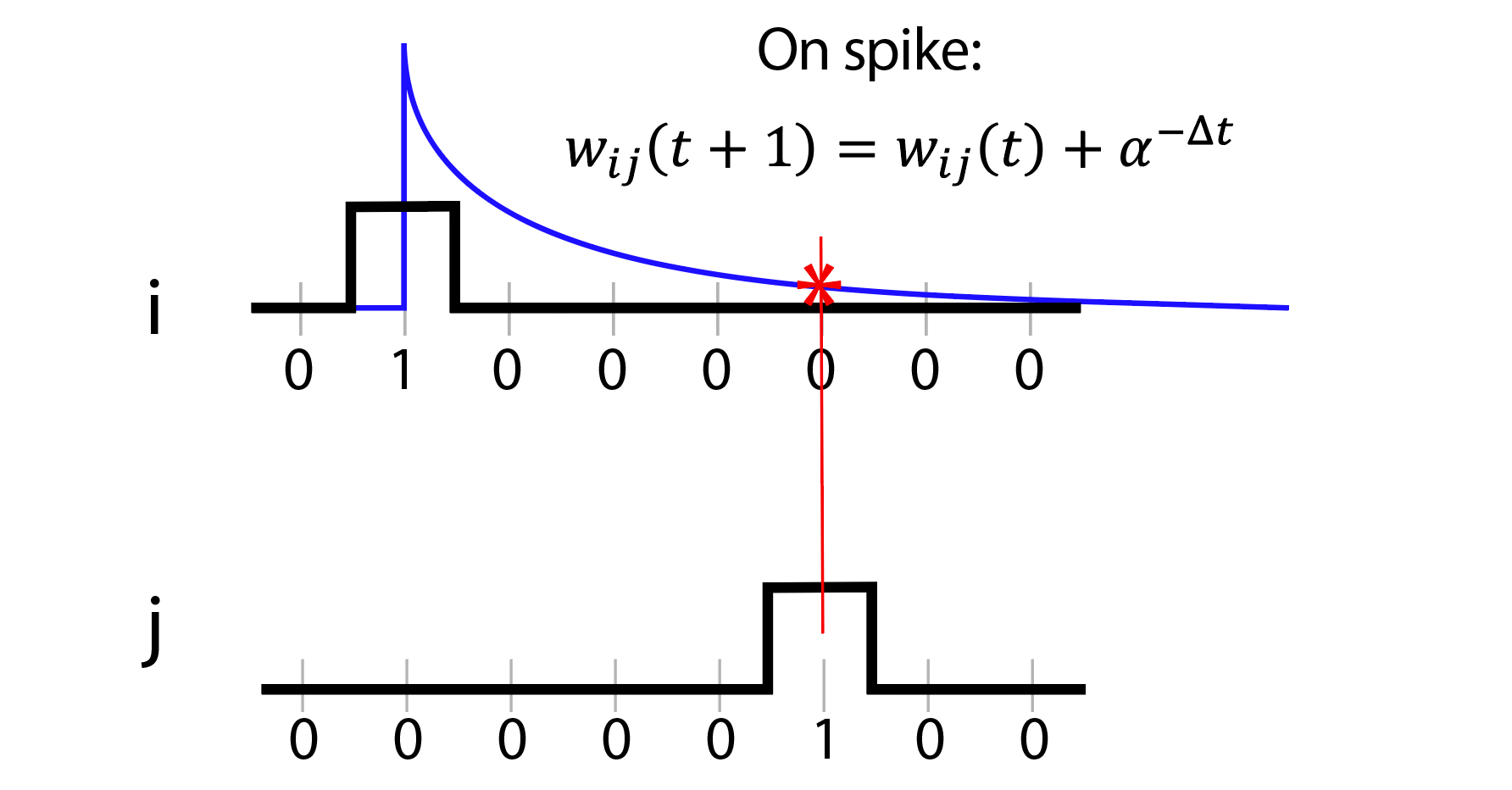
Spike Softening is well-suited for real-time application: it is purely causal and depends only on the previous sample at each time step, giving it a reduced memory footprint and better scaling compared to the state of the art. Its simplistic nature also enables straightforward implementation in highly parallel computing environments such as FPGAs or GPUs.
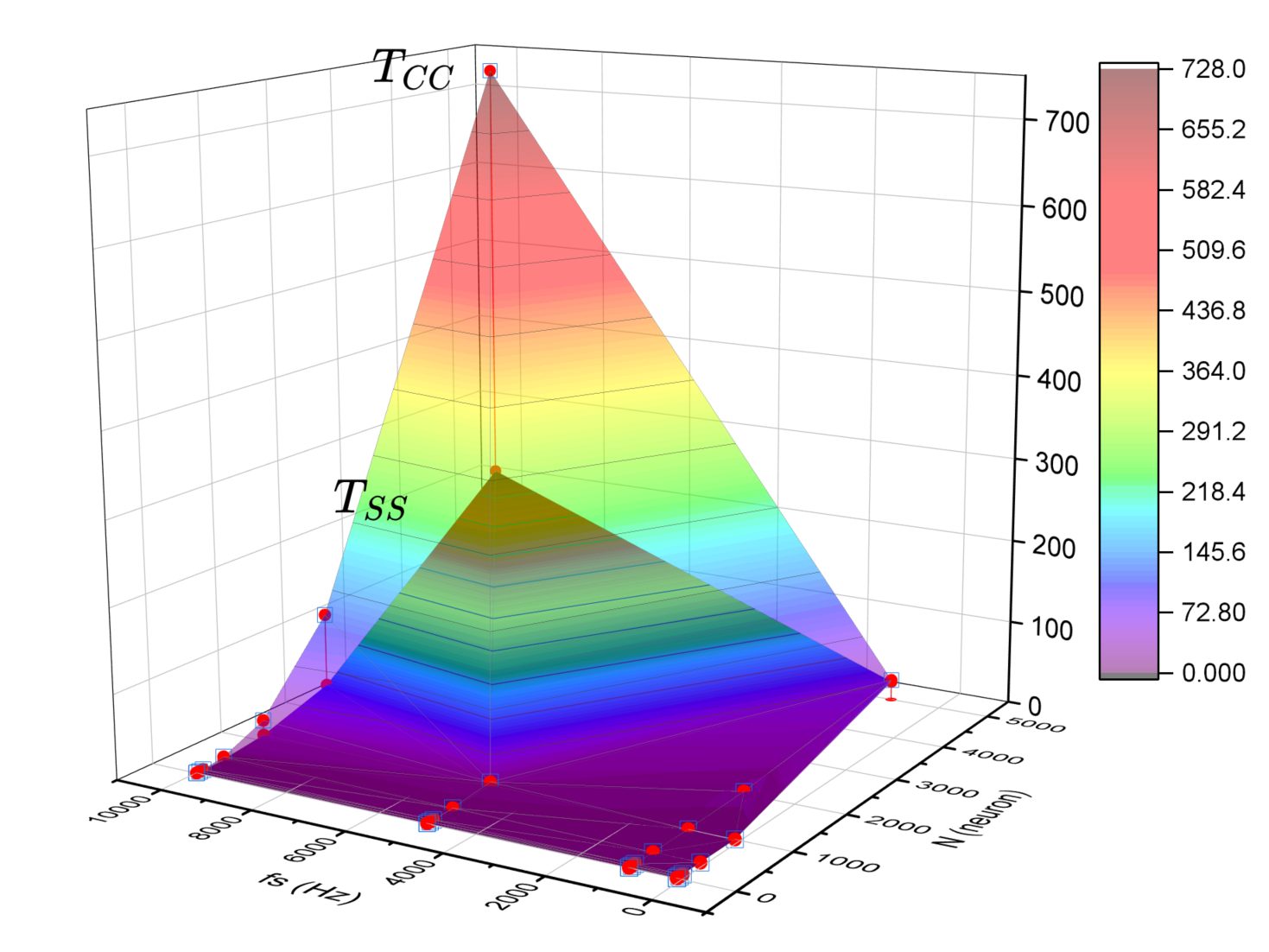
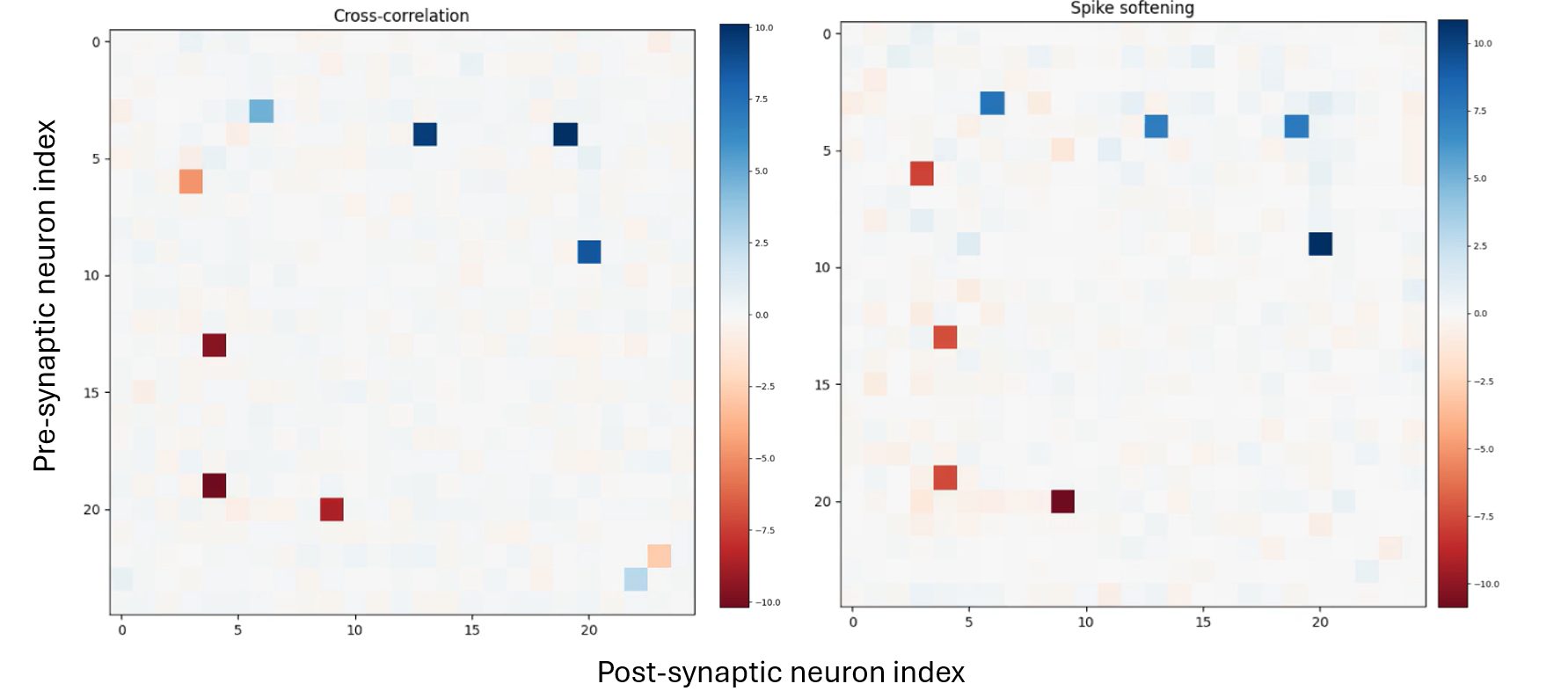
Quantitative validation was performed by comparing randomly generated weight matrices feeding a Stochastic Integrate-and-Fire (SIF) model against their cross-correlation and Spike Softening estimated counterparts; both algorithms performed at comparable accuracy scores, with faster computation on the latter. Experimental validation using a spike-sorted cat visual cortex recording dataset confirmed the same general network identification behaviour — the resulting connection matrices showed the same strong correlations in both methods.
Affiliated member
Claude Dahan — PhD Researcher
[email protected]