Swiss Data Science Center (SDSC) & Biomedical Informatics Group (ETHZ)
Earliest start date: January 2026
Background
Accurately estimating cardiac motion from cardiac Magnetic Resonance Imaging (MRI) data is a crucial step to evaluating cardiac function and detecting dysfunctions such as cardiomyopathy and infarction. The UK Biobank [1] offers an extensive dataset of cardiac MRI image volumes covering the ventricles of approximately 80,000 patients. These images are provided as temporal sequences (3D + T) that cover one heartbeat, enabling detailed tracking and modeling of myocardial tissue deformation. Previous work [2] has used a two-step pipeline consisting of landmark localization and shape registration to model myocardial motion, which is very expensive and constrained to predefined anatomical priors.
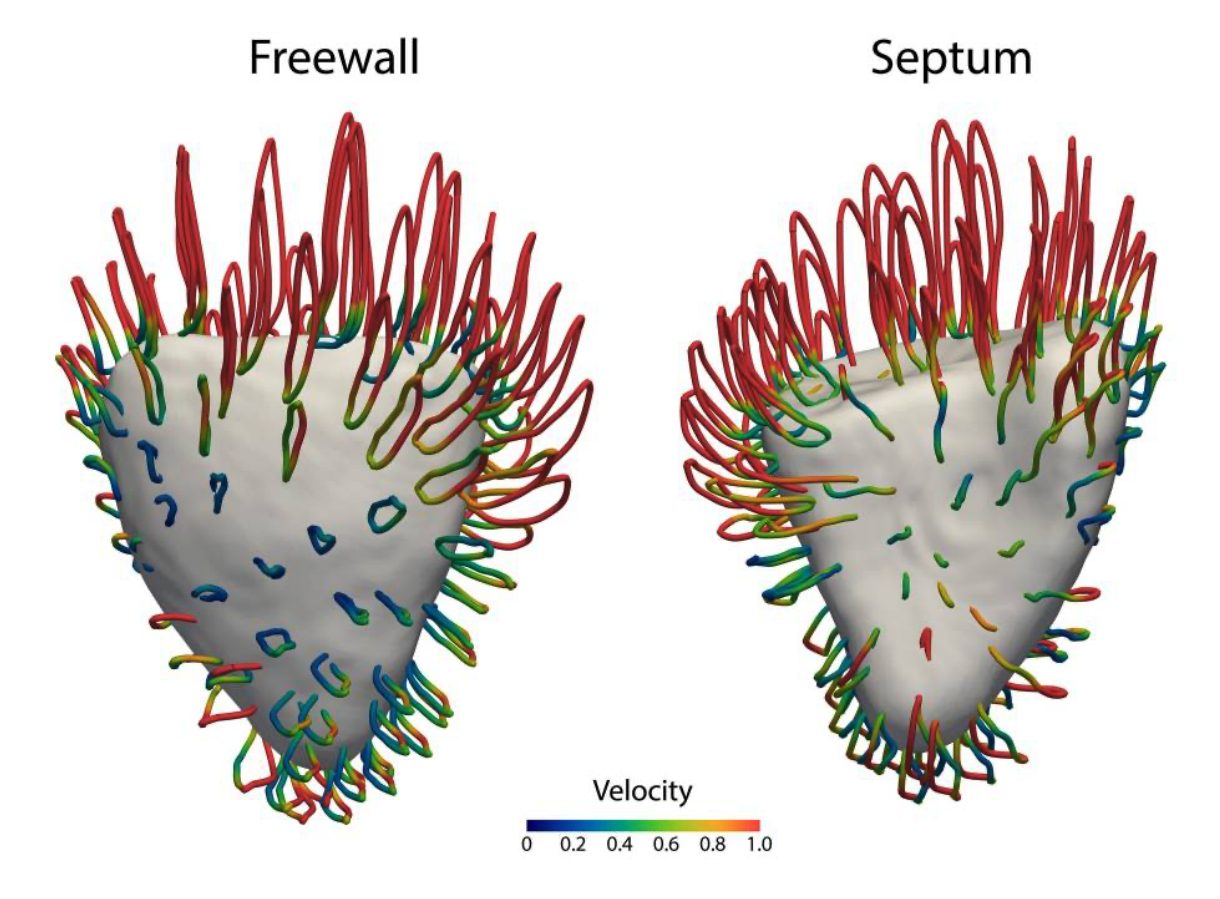
The primary objective of this project is to develop a probabilistic, spatio-temporal model of cardiac deformation that combines Gaussian primitives [3] with state-space Gaussian process (GP) dynamics. The aim is to achieve a physically interpretable, data-efficient, and uncertaintyaware description of myocardial motion throughout the cardiac cycle, directly from 3D + T MRI sequences.
Specific Objectives
- Represent myocardial tissue with Gaussian primitives. Each Gaussian primitive will describe a local tissue parcel in 3D space by a mean position, covariance shape, and intensity. This compact representation provides a continuous and differentiable description of myocardial geometry and texture.
- Model temporal deformations with state-space Gaussian processes. The temporal evolution of each primitive’s degrees of freedom—translation, rotation (via angular velocity), and local stretch—will be modeled using Markovian GP priors in state-space form. This approach allows the deformation field to be expressed as a system of stochastic differential equations (SDEs), enabling Kalman filtering and smoothing for efficient inference across time while maintaining continuity and temporal smoothness.
- Enhancing flexibility and lowering dimensionality using a latent GP. Directly modeling the primitives’ parameters with a GP can limit model expressiveness and increase computational cost. To overcome these challenges, a promising approach is to employ a latent GP that adaptively encodes memory, efficiently integrating information from previous views. This strategy is inspired by the method presented in [4].
- Validate on dynamic cardiac MRI. The resulting model will be evaluated on 3D + T cardiac MRI datasets, assessing its ability to capture realistic and smooth deformation patterns.
Additional Information
- What will you learn? The project will allow the student to gain hands-on experience with Gaussian splatting, spatio-temporal Gaussian processes, and probabilistic modelling in general.
- Requirements: Basic knowledge of Gaussian processes, good Python and PyTorch skills.
- Supervisors: – David Brüggemann ([email protected]) – Quentin Duchemin ([email protected]) – Olga Demler ([email protected])
- Publication? Possible if the project goes well.
- Interested? Contact us via email.
References
[1] Petersen, S.E., Matthews, P.M., Francis, J.M., Robson, M.D., et al.: Uk biobank’s cardiovascular magnetic resonance protocol. Journal of cardiovascular magnetic resonance 18, 8 (2016)
[2] Bello, G.A., Dawes, T.J., Duan, J., Biffi, C., et al.: Deep-learning cardiac motion analysis for human survival prediction. Nature machine intelligence 1, 95–104 (2019)
[3] Kerbl, B., Kopanas, G., Leimk¨uhler, T., Drettakis, G.: 3d gaussian splatting for real-time radiance field rendering. ACM Trans Graph 42, 139–1 (2023)
[4] Hou, Y., Kannala, J., Solin, A.: Multi-view stereo by temporal nonparametric fusion. In: Proceedings of the IEEE/CVF International Conference on Computer Vision, 2651–2660 (2019)